Research
MAQC Consortium 2011–2014 (read more)
FDA SEQC, Nature Biotechnology (read more)
Bioinformatics (read more)
FEBS Journal (read more)
Gerontology (read more)
In view of the extremely large number of potential probe sequences to consider, many approaches to microarray design use greedy search or filtering instead of true optimization across the parameter space. Similarly, heuristic shortcuts are employed in lieu of more elaborate thermodynamic models of the probe binding process.
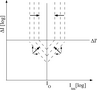 In Leparc et al. (2009), we have shown the improvements
to specificity, sensitivity, and array uniformity that can be
achieved by non-greedy set-based optimization of microarray
probe design. Furthermore, we have introduced an integrated
quantitative model of the effects of labelling,
cross-hybridization, and other probe and target structures
competing for hybridization.
In Leparc et al. (2009), we have shown the improvements
to specificity, sensitivity, and array uniformity that can be
achieved by non-greedy set-based optimization of microarray
probe design. Furthermore, we have introduced an integrated
quantitative model of the effects of labelling,
cross-hybridization, and other probe and target structures
competing for hybridization.
In Mückstein et al. (2010), we have shown the
benefits of an
improved model for microarray hybridization. Remarkably,
specific and unspecific hybridization were apparently
driven by different energetic contributions: For
unspecific hybridization,
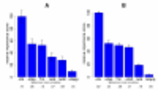 the probe–target melting temperature
Tm was the best predictor of signal
variation. For specific hybridization, however,
the effective interaction energy that also considered
alternative competing conformations was twice as
powerful a predictor of probe signal variation,
highlighting the importance of secondary structures in
the probe and target molecules.
the probe–target melting temperature
Tm was the best predictor of signal
variation. For specific hybridization, however,
the effective interaction energy that also considered
alternative competing conformations was twice as
powerful a predictor of probe signal variation,
highlighting the importance of secondary structures in
the probe and target molecules.
References
- Kreil DP, Russell RR, Russell S (2006) Microarray oligonucleotide probes. Methods Enzymol. 410, 73–98. (preprint on request | Supplement)
- Leparc GG, Tüchler T, Striedner G, Bayer K, Sykacek P, Hofacker IL, Kreil DP (2009) Model-based probe set optimization for high-performance microarrays. Nucleic Acids Res 37, e18. (read more | Supplement)
- Mueckstein U, Leparc GG, Posekany A, Hofacker I, Kreil DP (2010) Hybridization thermodynamics of NimbleGen microarrays. BMC Bioinformatics 11, 35. (read more)
Return to group home or publications