Research
MAQC Consortium 2011–2014 (read more)
FDA SEQC, Nature Biotechnology (read more)
Bioinformatics (read more)
FEBS Journal (read more)
Gerontology (read more)
Microarray hybridization thermodynamics
While microarrays are the predominant method for gene expression profiling, understanding the variation of measurement signals is still an area of active research. Signals are probe sequence dependent, and both the interpretation of measurements and the design of microarrays need to take this into account.
In view of the extremely large number of potential probe
sequences to consider, many approaches to microarray design
employ heuristic shortcuts in lieu of more
elaborate thermodynamic models of the probe binding
process. We here demonstrate the benefits of an
improved model for microarray hybridization and assess
the relative contributions of the probe-target binding
strength and various competing structures. Remarkably,
specific and unspecific hybridization were apparently
driven by different energetic contributions: For
unspecific hybridization,
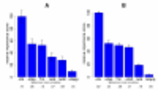 the probe–target melting temperature
Tm was the best predictor of signal
variation. For specific hybridization, however,
the effective interaction energy that also considered
alternative competing conformations was twice as
powerful a predictor of probe signal variation,
highlighting the importance of secondary structures in
the probe and target molecules.
the probe–target melting temperature
Tm was the best predictor of signal
variation. For specific hybridization, however,
the effective interaction energy that also considered
alternative competing conformations was twice as
powerful a predictor of probe signal variation,
highlighting the importance of secondary structures in
the probe and target molecules.
Reference
Mueckstein U, Leparc GG, Posekany A, Hofacker I, Kreil DP (2010) Hybridization thermodynamics of NimbleGen microarrays. BMC Bioinformatics 11, 35. (read more)
Return to group home or publications